My name's Eric. Welcome to my blog. 😊
A guide to productive game development in Rust when your machine can't keep up.
What better way to get better at a programming language than to write a game
in it? It's fun, creative, and complex enough to get you below the surface
level language features. Rust makes me think, and (unrelated to the rest of
this post but) Polars alone is enough of a
reason for an MLE to get involved.
So - Rust, games.
On the search for a framework,
Bevy stood out.
From their homepage, Bevy is a "refreshingly simple data-driven game engine
built in Rust". It uses native structs and has a mental model that is pretty
easy to "get dangerous" with.
Clean, crisp, awesome!
But not so fast.
My ancient macbook can't run any of even the simplest
examples from Bevy's extensive
list. Great sad. After a few months ruminating on this issue, trying
other projects here and there to salve the Bevy-shaped ache in my brain, this
time around I found some
success.
By some miracle (i.e. with lots of help from blog posts by those more skilled
than I,) I found a productive remote development workflow. With that ethos at
heart, I'll leave this attempt at a guide in hopes that it helps at least one
other interested tinkerer.
On with the tech!
See the related GitHub issues in the sources below for details on the shader translation error necessitating a workaround.
Game dev's workflow requirements are not as straightforward as other development: game development requires graphics, not just a terminal.
⏱about 1 hour, assuming you have an AWS account already.
💵less than $0.25; hourly t2.large was $0.1 at time of writing.
🏎
I’ve found it’s ok, but you could go for an xlarge or c family and be happier.
I put 64GiB EBS on it.
All other default options are ok, just make sure you have SSH access in the SG.
🍳
In corresponding order to the tl;dr:
Once everything is installed reboot the Ubuntu instance 📦♻️.
```sh
sudo apt update -y
sudo apt install -y xorg libxkbcommon-x11-dev
curl --proto '=https' --tlsv1.2 -sSf https://sh.rustup.rs | sh
sudo apt install -y build-essential pkg-config libasound2-dev libudev-dev
sudo apt install -y tigervnc-standalone-server tigervnc-xorg-extension tigervnc-viewer
sudo apt install -y lxde
```
Open an SSH connection to remote host and bind the 5901 port to localhost.
```sh
ssh ubuntu@ec2-[network-addr] -L 5901:127.0.0.1:5901
```
First, you will need to set a password.
```sh
vncpasswd
vncserver -xstartup /usr/bin/lxsession
```
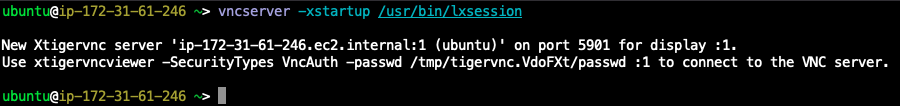
You can see the vnc servers with`vncserver -list`
and kill with`vncserver -kill :1`
Go ahead and leave this terminal with the SSH connection open.📡🫀
Reproduced with modification from the bevy example list:
```sh
git clone https://github.com/bevyengine/bevy
cd bevy
cargo run --example bloom_3d
```
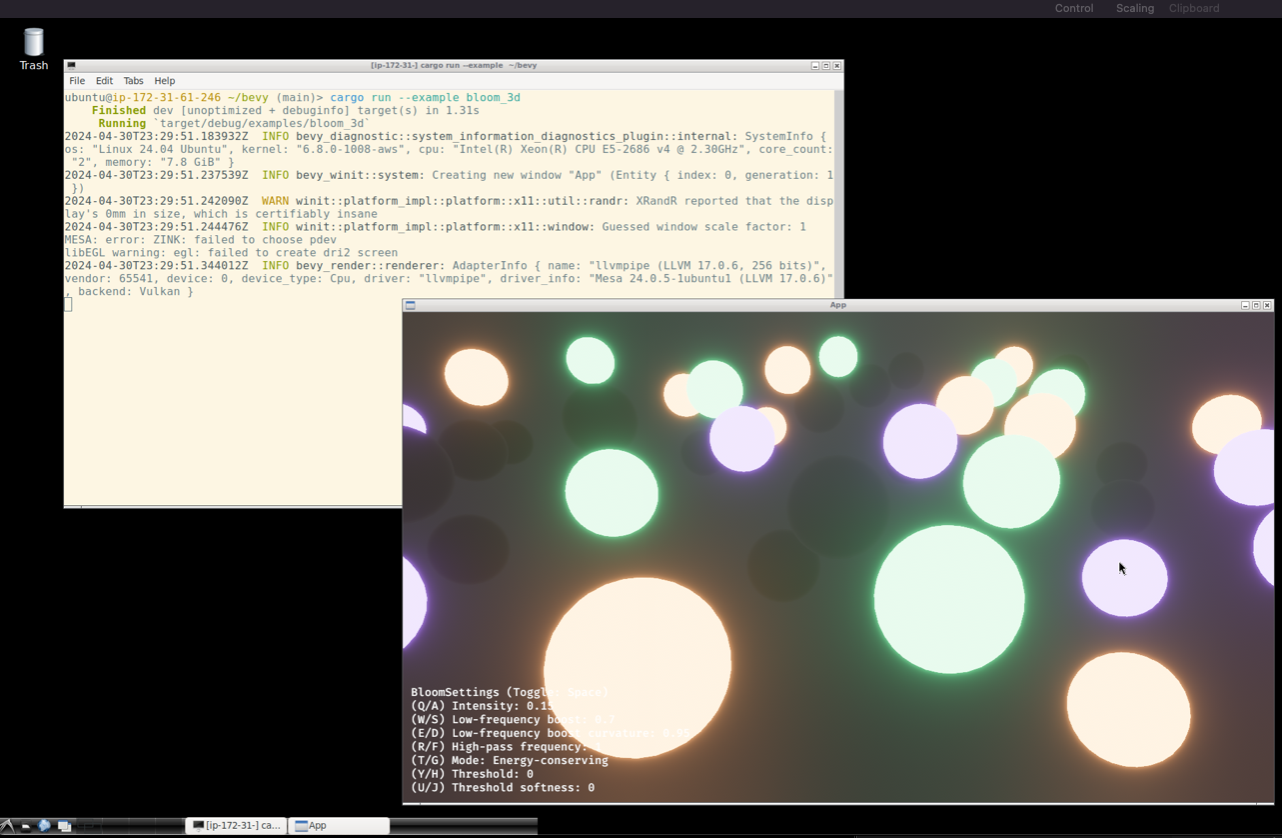
🏗🍭
In no particular order - hopefully some wary soul finds a clue here!
cargo run --target x86_64-apple-darwin --example bloom_2d
```Makefile
build-wasm:
@echo "🏎 Build start."
@date
@echo "📦 Compiling Rust code..."
cargo build \
--profile wasm-release \
--timings \
--target wasm32-unknown-unknown
@echo "🏗 Generating WASM..."
wasm-bindgen --out-name waste \
--out-dir web/target/wasm \
--target web target/wasm32-unknown-unknown/wasm-release/waste.wasm
@echo "🏁 Build complete."
@date
```
If you're interested, I've got a couple other posts.
License: MIT; Author: Eric Kalosa-Kenyon (2024); Find me on GitHub;